What are the Laws of Biology?
The reductionist perspective on biology is that it all boils down to physics eventually. That anything that is happening in a living organism can be fully accounted for by an explanation at the level of matter in motion – atoms and molecules moving, exerting forces on each other, bumping into each other, exchanging energy with each other. And, from one vantage point, that is absolutely true – there’s no magic in there, no mystical vital essence – it’s clearly all physical stuff controlled by physical laws.
But that perspective does not provide a
sufficient explanation of life. While living things obey the laws of physics,
one cannot deduce either their existence or their behaviour from those laws
alone. There are some other factors at work – higher-order principles of design
and architecture of complex systems, especially ones that are either designed
or evolved to produce purposeful
behaviour. Living systems are for
something – ultimately, they are for replicating themselves, but they have
lots of subsystems for the various functions required to achieve that goal. (If
“for something” sounds too anthropomorphic or teleological, we can at least say
that they “do something”).
Much of biology is concerned with working
out the details of all those subsystems, but we rarely discuss the more
abstract principles by which they operate. We live down in the details and we
drag students down there with us. We may hope that general principles will
emerge from these studies, and to a certain extent they do. But it feels like
we are often groping for the bigger picture – always trying to build it up from
the components and processes we happen to have identified in some specific
area, rather than approaching it in any principled fashion or basing it on any
more general foundation.
So, what are these principles? Do they even
exist? Can we say anything general about how life works? Is there any
theoretical framework to guide the interpretation of all these details?
Well, of course, the very bedrock of
biology is the theory of evolution by natural selection. That is essentially a
simple algorithm: take a population of individuals, select the fittest (by
whatever criteria are relevant) and allow them to breed, add more variation in
the process, and repeat. And repeat. And repeat. The important thing about this
process is it builds functionality from randomness by incorporating a
ratchet-like mechanism. Every generation keeps the good (random) stuff from the
last one and builds on it. In this way, evolution progressively incorporates
design into living things – not through a conscious, forward-looking process,
but retrospectively, by keeping the designs that work (for whatever the
organism needs to do to survive and reproduce) and then allowing a search for
further improvements.
But that search space is not infinite – or
at least, only a very small subsection of the possible search space is actually
explored. I often use a quote from computer scientist Gerald Weinberg, who said
that: “Things are the way they are because they got that way”. It nicely
captures the idea that evolution is a series of frozen accidents and that
understanding the way living systems are put together requires an evolutionary
perspective. That’s true, but it misses a crucial point: sometimes things are
the way they are because that’s the only way that works.
Natural selection can explain how complex
and purposeful systems evolve but by itself it doesn’t explain why they are the
way they are, and not some other way. That comes down to engineering. If you
want a system to do X, there is usually a limited set of ways in which that can
be achieved. These often involve quite abstract principles that can be implemented
in all kinds of different systems, biological or designed.
Systems
principles
Systems biology is the study of those kinds
of principles in living organisms – the analysis of circuits and networks of
genes, proteins, or cells, from an engineering design perspective. This
approach shifts the focus from the flux of energy and matter to emphasise
instead the flow of information and
the computations that are performed on it, which enable a given circuit or
network to perform its function.
In any complex network with large numbers
of components there is an effectively infinite number of ways in which those
components could interact. This is obviously true at the global level, but even
when talking about just two or three components at a time, there are many
possible permutations for how they can affect each other. For a network of
three transcription factors, for example, A could activate B, but repress C; or
A could activate B and together they could repress C; C could be repressed by
either A OR B or only when both A AND B are present; C could feed back to
inactivate A, etc., etc. You can see there is a huge number of possible
arrangements.
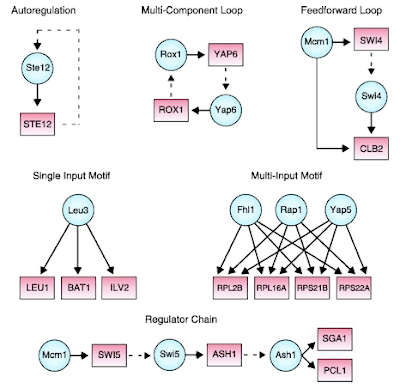
The core finding of systems biology is that
only a very small subset of possible network motifs is actually used and that
these motifs recur in all kinds of different systems, from transcriptional to
biochemical to neural networks. This is because only those arrangements of
interactions effectively perform some useful operation, which underlies some
necessary function at a cellular or organismal level. There are different
arrangements for input summation, input comparison, integration over time, high-pass
or low-pass filtering, negative auto-regulation, coincidence detection,
periodic oscillation, bistability, rapid onset response, rapid offset response,
turning a graded signal into a sharp pulse or boundary, and so on, and so on.
These are all familiar concepts and designs
in engineering and computing, with well-known properties. In living organisms
there is one other general property that the designs must satisfy: robustness.
They have to work with noisy components, at a scale that’s highly susceptible
to thermal noise and environmental perturbations. Of the subset of designs that
perform some operation, only a much smaller subset will do it robustly enough
to be useful in a living organism. That is, they can still perform their
particular functions in the face of noisy or fluctuating inputs or variation in
the number of components constituting the elements of the network itself.
These robust network motifs are the
computational primitives from which more sophisticated systems can be
assembled. When combined in specific ways they produce powerful systems for
integrating information, accumulating evidence and making decisions – for
example, to adopt one cell fate over another, to switch on a metabolic pathway,
to infer the existence of an object from sensory inputs, to take some action in
a certain situation.
A
conceptual framework for systems biology
Understanding how such systems operate can
be greatly advanced by incorporating principles from a wide range of fields,
including control theory or cybernetics, information theory, computation theory, thermodynamics, decision theory, game theory, network theory, and many
others. Though each of these is its own area, with its own scholarly
traditions, they can be combined into a broader schema. Writing in the 1960’s, Ludwig
von Bertalanffy – an embryologist and philosopher – recognised the conceptual
and explanatory power of an over-arching systems perspective, which he called
simply General System Theory.
Even this broad framework has limitations,
however, as does the modern field of Systems Biology. The focus on circuit
designs that mediate various types of information processing and computation is
certainly an apt way of approaching living systems, but it remains, perhaps,
too static, linear, and unidirectional.
To fully understand how living organisms
function, we need to go a little further, beyond a purely mechanistic
computational perspective. Because living organisms are essentially
goal-oriented, they are more than passive stimulus-response machines that
transform inputs into outputs. They are proactive agents that actively maintain
internal models of themselves and of the world and that accommodate to incoming information by updating those models and altering
their internal states in order to achieve their short- and long-term goals.
This means information is interpreted in
the context of the state of the entire cell, or organism, which includes a
record or memory of past states, as well as past decisions and outcomes. It is
not just a message that is propagated through the system – it means something to the receiver and that
meaning inheres not just in the message itself, but in the history and state of
the receiver (whether that is a protein, a cell, an ensemble of cells, or the
whole organism). The system is thus continuously in flux, with information
flowing “down” as well as “up”, through a constantly interacting hierarchy of
networks and sub-networks.
A mature science of biology should thus be
predicated on a philosophy more rooted in process than in fixed entities and
states. These processes of flux can be treated mathematically in complexitytheory, especially dynamical systems theory, and the study of self-organising
systems and emergence.
In addition, the field of semiotics (the
study of signs and symbols) provides a principled approach to consider meaning.
It emerged from linguistics, but the principles can be applied just as well to
any system where information is passed from one element to another, and where
the state and history of the receiver influence its interpretation of the
message.
In hierarchical systems, this perspective yields
an important insight – because messages are passed between levels, and because
this passing involves spatial or temporal filtering, many details are lost
along the way. Those details are, in fact, inconsequential. Multiple physical
states of a lower level can mean the same thing to a higher level, or when
integrated over a longer timeframe, even though the information content is
formally different. This means that the low-level laws of physics, while not
violated in any way, are not sufficient in themselves to explain the behaviour
of living organisms that process information in this way. It requires thinking
of causation in a more extended fashion, both spatially and temporally, not solely
based on the instantaneous locations and momentum of all the elementary
particles of a system.
A
new pedagogical approach in Biology
In the basic undergraduate Biology textbook
I have in my office there are no chapters or sections describing the kinds of principles
discussed above. There is no mention of
them at all, in fact. The words “system”, “network”, “computation” and
“information” do not even appear in the index. The same is true for the
textbooks on my shelf on Biochemistry, Molecular and Cell Biology,
Developmental Biology, Genetics, and even Neuroscience.
Each of these books is filled with detail
about how particular subsystems work and each of them is almost completely
lacking in any underpinning conceptual theory. Most of what we do in biology
and much of what we teach is describing what’s happening – not what a system is doing. We’re always trying to figure out how some particular system
works, without knowing anything about how systems work in general. Biology as a
whole, along with its sub-disciplines, is simply not taught from that
perspective.
This may be because it necessarily involves
mathematics and principles from physics, computing, and engineering, and many
biologists are not very comfortable with those fields, or even acutely math-phobic.
(I’m embarrassed to say my own mathematical skills have atrophied through
decades of neglect). Mainly for that reason, the areas of science that do deal
with these abstract principles – like Systems Biology or Computational
Neuroscience, or more generally relevant fields like Cybernetics or Complexity
Theory, are ironically seen as arcane specialties, rather than providing a
general conceptual foundation of Biology.
As we have learned more and more details in
more and more areas, we seem to have actually moved further and further away
from any kind of unifying framework. If anything, discussion of these kinds of
issues was more lively and probably more influential in the early and mid-1900s
when scientists were not so inundated by details. It was certainly easier at
that time to be a true polymath and to bring to bear on biological questions
principles discovered first in physics, computing, economics, or other areas.
But now we’re drowning in data. We need to
educate a new breed of biologists who are equipped to deal with it. I don’t
mean just technically proficient in moving it around and feeding it into black
box machine-learning algorithms in the hope of detecting some statistical
patterns in it. And I don’t necessarily mean expert in all the complicated
mathematics underlying all the areas mentioned above. I do mean equipped at
least with the right conceptual and philosophical framework to really
understand how living systems work.
How to get to that point is the challenge,
but one I think we should be thinking about.
Resources
Science and the Modern World. Alfred North
Whitehead, 1925.
What is Life? Erwin Schrodinger, 1943.
Cybernetics. Or Control and Communication in the Animal and the Machine. Norbert Wiener, 1948.
Cybernetics. Or Control and Communication in the Animal and the Machine. Norbert Wiener, 1948.
The Strategy of the Genes. Conrad
Waddington, 1957.
The Computer and the Brain. John von
Neumann, 1958.
General System Theory. Foundation, Theory, Applications. Ludwig von Bertalanffy,
1969.
Gödel, Escher, Bach. An Eternal Golden Braid. Douglas Hofstadter,
1979.
The Extended Phenotype. Richard Dawkins.
1982.
Order out of Chaos. Man’s New Dialogue with
Nature. Ilya Prigogine and Isabelle Stengers, 1984.
Endless Forms Most Beautiful. Sean Carroll,
2005.
Robustness and Evolvability in Living Systems.
Andreas Wagner, 2007.
Complexity: A Guided Tour. Melanie
Mitchell, 2009.
The Information. A History, A Theory, A Flood. James Gleick, 2011.
Cells to Civilisations. The Principles of Change that Shape Life. Enrico Coen, 2015.



I have been obsessing about these very same questions myself.
ReplyDeleteI think these is a great need and an opportunity for a sketch of "the large picture" that would be based on scientific understanding of reality, and be useful to scientists -- something that philosophy could have done, but presently have not.
As Feynman says in this interview, everything is atoms merely "doing their thing":
https://youtu.be/m6BqcO-U4R0
But what is very often inexplicably forgotten, is that microscopic atomic picture is not "just physics." Crucially, *the arrangement* of the atoms determines how the elementary, fundamental actions add together or cancel each other.
Thus, the actual microscopic description is:
(Full structure/state of the system) x (fundamental laws that make it go)
This simply means that the history of the evolving (state/structure) does not follow from fundamental laws of physics -- it just as much depends on the (state/structure) itself. (Which determines how the elementary, fundamental actions add together or cancel each other and in doing so modify the state/structure -- creating the dynamics of the system.)
Without describing the (state/structure), fundamental physics does not describe *our* world, but merely all possible worlds consistent with the laws of physics.
(Paradoxically, the set of "all that can be" is a much less complex thing than any one of its members: Think of the library of all strings -- it can be trivially specified, and contains almost no information. The act of choosing one "book" is what creates the information contained in it.)
Thus, "reduction" to the laws of physics is not possible. We must include the history of the evolving (structure/state) as it actually unfolds. (Or at least the key stepping stones of this history which pin it to a particular course of development.)
Of course, nature existed long before life and humans came to be.
Not always, but in many important cases the dynamics creates structures that are "lumpy" and action that is confined to narrow "channels." Atoms, planets, atmospheric circulation, organisms all provide examples of such lumps and channels. The "geometry of action" in nature is the networks of chemical reactions, the lace of rivers on the planet, etc, etc:
https://twitter.com/generuso/status/894738764664360960
The lumpiness and channeling mean that the evolution of state/structure is confined to a tiny manifold within its full microscopic state space. This manifold represents the macroscopic structure of reality, and comes with its own rules of action.
Discerning the fundamental principles of the structure of reality, in my view is the necessary foundation to sketching the large picture of life, evolution and cognition.
Thanks for your comments and insights. I completely agree. The issue is to try and define and understand the rules of action or underlying organisational principles of living things - what makes them the way they are and not some other way? What gives them the emergent properties of life, including agency and ultimately consciousness? These are philosophical questions but they have clear practical implications in terms of us making real progress towards understanding living things well enough to predict things like the effects of mutations in a given gene, the effect of a new drug and how the system might respond to it, the emergent pathology in neurodevelopmental disorders and the ultimate reasons for the patterns of psychopathology observed, etc, etc.
DeleteAnother lovely swat at complexity- thanks much, Kevin, especially for the references and the graphic, which has veered me unduly from my normal life. Sometimes it's a great feeling to feel left behind, at least when one is handed maps and hypertexts.
Delete> define and understand the rules of action or underlying organisational principles of living things - what makes them the way they are and not some other way? What gives them the emergent properties of life, including agency and ultimately consciousness? These are philosophical questions
If we might turn this on its head a moment– one should argue occasionally that they're not philosophical questions at all, but practical ones that philosophers might build upon fruitfully, if they pay attention and behave themselves. What I find exciting about the evaluation of biological systems is the ability to virtually prohibit philosophical lines of inquiry that conflict with findings (especially pop philosophy), and to bring to light unimagined philosophical dimensionality through leveraging the long aeons of hit-and-miss that got we-the-living so incredibly successful and complex. Building up from what actually happens in our living systems will lay bare useless, less useful, and more useful assumptions lodged in the layers and layers of abstractions and levels of analysis that make up the notions of will and consciousness.
Subconsciously, we all cringe at the notion of being soft machines, I suspect as a by-product of very basic selection optimizations, the same ones that encouraged both religiosity and practical diversity within populations. This reluctance to "name" ourselves as machines makes us dichotomize unduly between non-human aspects of behaviour, will and consciousness, and our own versions. That dichotomizing has the side effect of fetishizing the will and consciousness, instead of seeing them as complex/dynamical extensions of first principles. In contrast, building to these abstractions from biology allows us to see them as natural extensions of robustness and fundamental circuitry, and to learn how the natural world dictates its working philosophies. The is of 'is and ought' has been incredibly successful within biology, and yet we look constantly beyond ourselves as writ to find coherence and meaning. Denying ourselves as amazingly successful machinery propels us to feel instantiated by ties to deity instead, or worse, it justifies tragic, narrowly-defined individual "successes" to establish meaning. The results of such bad modeling, in a much more interdependent world, are increasingly toxic and unworkable.
The power and joy inherent in the cybernetic-cum-social nature of life will be one of the great works of the soft and hard sciences intermixing in the coming decades. Is will get its say in building up to our oughts, just as the pragmatists have insisted should happen since the end of the nineteenth century, when God went behind a paywall.
Thanks Scott - I think you're exactly right, actually. If we approach these questions in a principled, scientific fashion, we can define things like the emergence of agency and even eventually of consciousness and will in a way that sharply delimits the otherwise potentially unbounded and never-ending philosophical discussions of these topics.
DeleteIt amazes me how much of the philosophical literature on things like free will is uninformed by our growing understanding of the actual mechanisms of decision-making - not just at the neural level, where we are making real progress, but in terms of the types of computations and operations that are happening, which you can see even at low levels like bacterial chemotaxis, or turning on the lac operon. There are common underlying principles of exploration vs exploitation, integration and accumulation of evidence, estimation of confidence, internal measures of utility, etc., that can be seen to apply across all these kinds of decisons, from the least to the most sophisticated.
And, of course, there are many people working to draw out these principles, but somehow those efforts remain outside the mainstream in Biology - certainly in how we teach students. Time to try and do better!
Couple of things.
ReplyDelete> Now we're drowning in data.
This isn't exactly unfair to say, but it does discount the much larger challenge that polymaths have nowadays, now that we know more, to establish system principles. There's a natural evolution in any project at any scale to know very little, to systematize it avidly, reject that in favor of reality, and then to simplify again in a more sophisticated and accurate fashion. You're stuck on that last step, and doing a helluva lot better with it than the physicists are.
And look at how incredibly difficult and interdisciplinary the robust science of systematizing is, in a world often dominated by nonlinear dynamics of various kinds. Pick any two scholars on that graphic you forwarded, and ask a physicist, or a psychologist, or even a biologist/statistician to understand their work without extraordinary labour, and they will fail. We are fighting not only deep cognitive blocks on interdisciplinary thinking, but also the difficult essence of reality's complex nature, that has us looking into the maelstrom for the first time, typically without the basic education to do so usefully.
Unrelated- the notion of robustness is tied to the notion of will, in the broad sense of slimming down possibilities to get at something that works. In selection, the proxy for the will is the rejection of unworkable systems longitudinally, just as a hunter 'rejects' hill and dale on the way to success. I like that connection, a lot; it makes the will a natural extension of robustness, which makes the notion of it simpler and more powerful to me.
Thanks Scott - this problem of seeing across disciplines is exactly the one I think we need to address, fundamentally, in early Biology education. How can we ever expect students to understand perspectives outside the narrow disciplinary areas they will inevitably end up in if we don't even make any effort to make them aware of the deep conceptual principles that play across all areas? Of course, most Biology lecturers weren't trained that way themselves. Even within disciplines, the degree of specialisation required to make progress (apparent progress at least) in research is extreme. Because that is what is valued in grant applications, promotions, etc., generalists are in short supply. (And true polymaths are a thing of the past, I think - there's just too much to know!)
DeleteAs a PhD student in Biology Education (teacher training) I totally agree with your statement about these perspectives in how we teach biology. I.e. complexity science is far from current classrooms and we need to change that. Our recent poster presented at the Cultural Evolution Society highlights a Design-Based Research approach to moving teacher training and classroom activities in exactly this direction.
ReplyDeletehttps://www.researchgate.net/publication/319644966_Cultural_Evolution_in_the_Biology_Classroom_A_Design-Based_Research_Model_in_Education_for_Sustainable_Development
Thanks Dustin - looks really interesting.
Delete"The core finding of systems biology is that only a very small subset of possible network motifs is actually used and that these motifs recur in all kinds of different systems, from transcriptional to biochemical to neural networks. This is because only those arrangements of interactions effectively perform some useful operation, which underlies some necessary function at a cellular or organismal level."
ReplyDeleteWell maybe, but my understanding is that for almost all systems we don't have enough information to fully describe how the system really works. Most scientific studies seem to give a pretty indirect and approximate view of one aspect of how the system functions. (And calling anything a system is probably a bit generous because all biological processes are so massively interactive that isolating one part of the whole is already a simplification.) With these small fragments of approximate knowledge it is possible to build a simplified, human-friendly narrative that puts together a group of organic compounds as if they functioned as some kind of biological printed circuit. But it's not really like that, and we know it isn't. The more one moves towards systems explanations the more one's science gets further from chemistry and closer to economics. It's an interesting perspective but I'm not convinced it will give us a clearer understanding of what's going on. And I think there is a risk that one will gain an illusory insight rather than a real one.
"This means information is interpreted in the context of the state of the entire cell, or organism, which includes a record or memory of past states, as well as past decisions and outcomes. It is not just a message that is propagated through the system – it means something to the receiver and that meaning inheres not just in the message itself, but in the history and state of the receiver (whether that is a protein, a cell, an ensemble of cells, or the whole organism)."
I'm afraid I don't really buy this. I'm not really sure that all living organisms have a model of themselves and their environment. I certainly baulk at the notion of most cells having a memory in any meaningful sense, and certainly not of their "past decisions and outcomes". I'd also need a bit of convincing that the history of a protein was significant, rather than its state.
I feel that this article describes the way things might be but that in fact we are very far from having enough real world data to say how things actually are. It seems that the more biology we discover the more complicated things get. I expect we'll be able to describe systems which provide a summary of the empirical events we observe, such as mounting an immune response, and that we will gain insights into microscopic aspects of those systems. But I'm doubtful that we'll get to a stage where we have strong theoretical understanding of the main features linking the microscopic and macroscopic phenomena. We'll always have a range of theories which seem plausible and we'll choose between them based on which most accurately fits the observed data. My understanding is that we're not even in a position to understand how a trained neural network achieves its function even though we can examine every aspect of it and even though it has a very simple architecture.
I suppose I think that the task of properly understanding biological processes is hopeless. So that instead we work pragmatically to try to understand little bits about the important bits which might have some useful applications. I think if we look at things mainly as systems we may be not very good at understanding or predicting how and why they stop working properly.
Thanks Dave. I think your skepticism is somewhat justified but I am more optimistic than you. I am not advocating abandoning experimentation on isolated components - not at all. What I am suggesting is that the design and interpretation of such experiments will be greatly advanced by considering the more abstract systems principles involved - by trying to figure out what the system is *doing*, not just describing what is happening.
DeleteIf the goal is to gain some predictive power about how and why systems fail, I would say the purely reductionist approach itself has been an abject failure. For example, we have a handful of new drugs every year, some very specifically designed to target particular molecules. But for each success, we have hundreds that fail in pre-clinical or clinical trials, often due to some unexpected systems-level effects.
For me, the systems perspective provides a complement to the study of experimentally isolated components. These are not actually isolated, as you point out, but it is possible to discern modules and motifs that do specific jobs, with specific dynamics and input-output relationships, and then to build larger systems from those components (where higher level "chunk" information from lower levels, not caring about the details).
Of course, it is possible to make nice toy mathematical models of complex systems and fool ourselves into thinking we understand them. But such models should make testable predictions and can be refined or rejected based on new experimental findings. That cycle of data -> theory -> prediction -> data -> new theory is certainly more powerful than the one-way system most of us are currently engaged in: data -> more data -> more data -> umm... more data?
I absolutely agree with Scott! These are practical questions that require answers based on a scientific view of reality.
ReplyDeleteEven some philosophers (for example Dennett, referring to the earlier word by W.Sellars) agree that the "manifest image of reality" on which everyday common sense and much of philosophy are based are not adequate for answering most of the serious questions in cognitive science.
Here is a nice lecture on this topic: "Professor Daniel C Dennett: 'Ontology, science, and the evolution of the manifest image'" https://www.youtube.com/watch?v=GcVKxeKFCHE
The questions of agency/will/consciousness are particularly heavily laden with antique, ingrained prejudices and naive intuitions.
Much too often, philosophers desire to jump in one leap from fundamental physics to consciousness, because our intuition tells us that consciousness must be fundamental, elementary.
But not long ago, people felt the same way about "life-force." It had been presumed to be fundamental, elementary ether that could make a lump of dirt into a live mouse.
Then it turned out to be something else entirely! A microscopic cell turned out to be a factory filled with a vast amount of intricate machinery! It was not simple at all! It was actually mindbogglingly complex, and that is what made the end result so unlike the elementary ingredients from which it is ultimately constructed.
Today we have plenty of examples of complex machinery which does things nothing short of magic. Does a monster in a video game look anything like the myriads of voltages in the computer chip that it "really is?" And this is "only" a machine with 10^10 elements!
The brain has 10^16 synapses not counting the rest of the good stuff. Do our everyday intuitions have any claim to be valid in predicting what machines of such complexity can or cannot do? Or why BRAIN ACTIVITY APPEARS TO ITSELF THE WAY IT DOES -- which is what consciousness really is?
Until we understand the functional architecture of brain machinery, it is very presumptuous to make philosophical claims as to the imaginary limitations of mechanistic explanations.
I agree. We need to build this conceptual framework from detailed data and experimentation, but we can do that in a more theoretically informed way than we have been doing (especially taking more abstract systems principles into account). Armchair philosophising will never get us there by itself - nature is not just stranger than we imagine, but stranger than we can imagine. On the other hand, just collecting more and more data and hoping these principles will spontaneously reveal themselves is also naive. The successes of methodological reductionism do not justify theoretical reductionism.
DeleteKevin,
ReplyDeleteRegarding your question: "Is there any theoretical framework to guide the interpretation of all these details?"
I have actually articulated exactly that. I have called it a new Intellectual Framework for science and for several years I have tried to get people to give it consideration. It remains unrefuted, and unacknowledged in the scientific community. Perhaps there is something amiss with THAT process?
I've also wrestled with this. Some answers may lie in this type of analysis: https://ombamltine.blogspot.com/2017/09/nonlinear-complex-phenomena-in-biology.html and the realization the complexity may arise out of iterated, more simple expressions.
ReplyDeleteJust a note to say that this very snazzy map (https://4.bp.blogspot.com/-ZAsaPSVsHno/WcuA8MtOFyI/AAAAAAAAA2I/uQqk2ZT9G0cLeUEeG1T-SbKhZL3vYI48gCLcBGAs/s1600/Complexity%2BTheory.png) is quite well documented as a terribly inaccurate, unscientific model which misses whole spaces, allocates concepts to the wrong people etc. It's great to get a general idea of some aspects, but please don't take it as gospel.
ReplyDeleteThank you for outlining the scope of this challenge, this is a fascinating subject to me. As someone who has been interested in trying to apply systems theory to biology for many years I have a question for you and for research scientists in any discipline within biology.
ReplyDeleteCould homeostasis (and/or allostasis) be considered to be in some sense a law of biology? If not a law then perhaps an organizing principle?
George F R Ellis, a mathematician and physicist, argues that homeostasis could be considered to be an example of top-down causation because it uses cybernetic feedback mechanisms to achieve a higher level process goal of maintaining relative equilibrium. So a macrostate process determines changes in microstates whether it occurs in a thermostat, a computer or in an organism.
Homeostatic process in organisms can be broken down into discrete mechanisms within subsystems. But is there measurable evidence of a higher level process goal for the whole body? Is body temperature a valid measure of whole body equilibrium with definable mechanisms? If so, could that be evidence of an organizing principle of how the body conforms to the second law of thermodynamics?
In terms of natural selection do new adaptations have to pass a homeostatic test? That is, can a new adaptation persist if it negatively affects the organism's ability to maintain homeostasis? Or conversely, does natural selection use the mechanisms of homeostasis to facilitate adaptations? For example, is the endocrine system a mechanism of selectively inducing disequilibrium within the organism as a means of motivating behavior to act and procure nutrients/sex/sleep etc? (This is my speculation that follows from Antonio Damasio's hypothesis that emotion/instinct/feeling are the collective motivating expressions of physiological homeostasis.)
Thanks for sharing.
ReplyDelete